Bayesian (and frequentist) re-analyses of published RCTs
These Shiny apps provide interactive Bayesian (and frequentist) re-analyses of published randomized controlled trials. They are part of the Assessing clinical trial interpretability research project funded by Heart & Stroke Canada.
Enter the 2×2 event counts from any RCT, choose an effect measure (relative risk or risk difference), and explore how different prior assumptions shape posterior conclusions.
Static entry image.
Bayesian RCT Re-analysis (Bayes only)
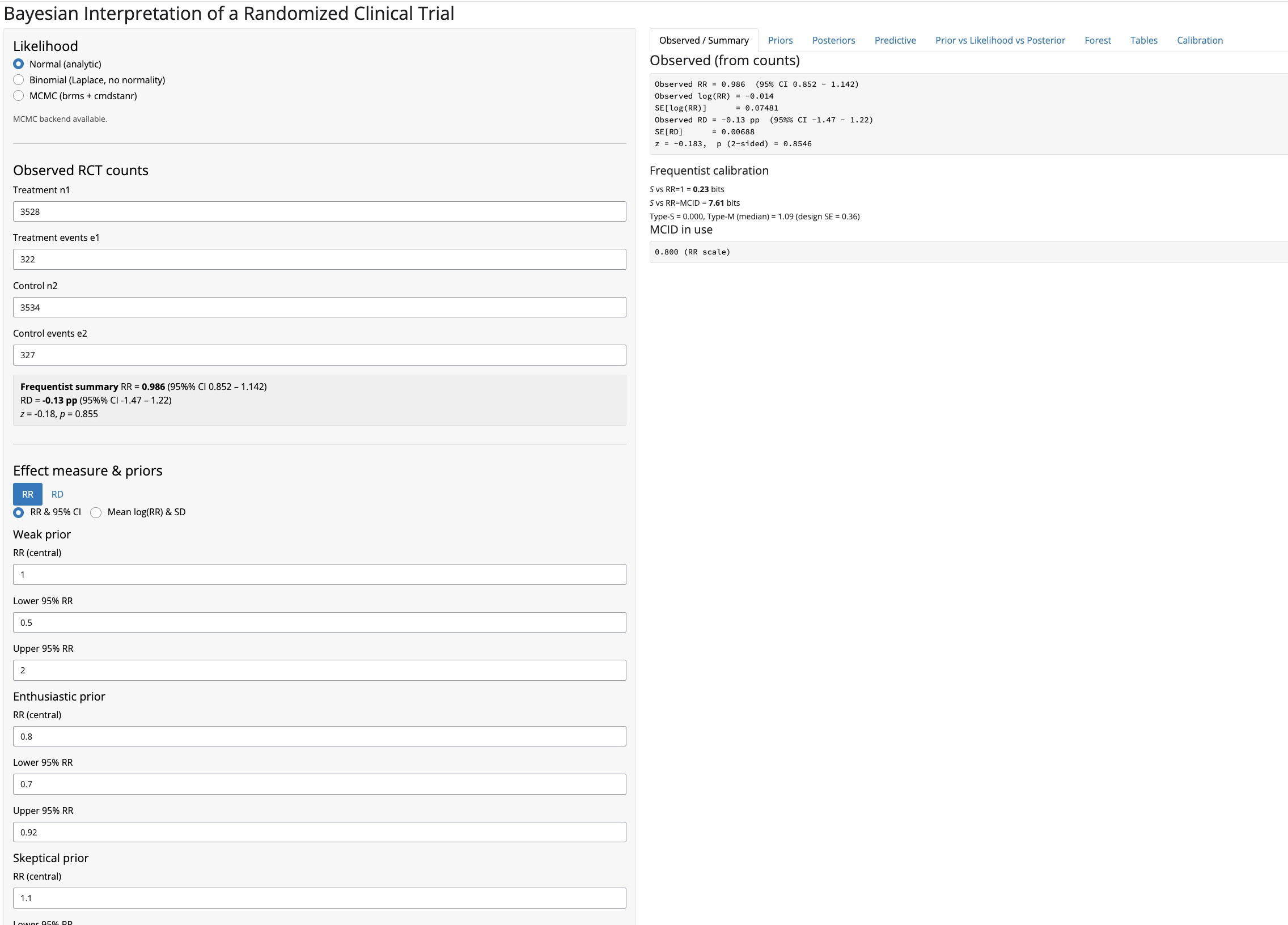
A focused Bayesian app with four priors — Weak, Enthusiastic, Skeptical, and Empirical Bayes (van Zwet et al. 2021/2025) — on either the relative risk (RR) or risk difference (RD) scale.
Features
- Three likelihood options: Normal (analytic), Binomial (Laplace), MCMC (brms)
- RR and RD effect measures via tabbed prior panels (priors entered in natural units)
- MCID-aware posterior summaries, predictive distributions, and forest plots
- Prior vs Likelihood vs Posterior facet plots
- CSV export of all summaries
Sample static output image.

Source code on GitHub
Full Bayesian + Frequentist Re-analysis
Everything in the Bayesian app, plus frequentist calibration metrics and an optional Design-informed prior.
Additional features
- Greenland S-values (vs RR = 1 and vs RR = MCID)
- Gelman–Carlin Type-S and Type-M error analysis
- Per-prior posterior calibration table (Type-S, Type-M, coverage, replication probability)
- Empirical-Bayes mixture component decomposition (Bk, B̄)
- Optional 5th Design-informed prior centred on the trial’s protocol target (RR mode)
Source code on GitHub
Clone the shiny_light repository and run either app in RStudio or from the command line:
shiny::runApp("app_bayes_alone.r")
shiny::runApp("app_full.r")Requires R ≥ 4.1 with shiny, ggplot2, dplyr, ggdist, tibble, bslib, and rlang. The MCMC backend additionally requires brms and cmdstanr (or rstan).
Use the shiny.io link